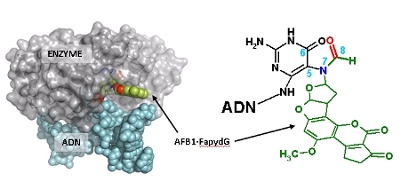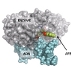
Combining chemical synthesis of derivatives DNA molecules, designed to mimic small and large lesions, with comparative structural analysis of Fpg bound to damaged DNA, scientist from CBM have established the molecular basis for formation of an unproductive but stable complex between Fpg and the bulky DNA lesion. They suggest that formation of such abortive complex inhibits DNA repair processes, thus explaining the long persistence of these lesions observed in vivo their associated cell deleterious effects. Selective inhibition of Fpg and hOgg1 by bulky DNA lesions and their synthetic derivatives can, for example, be used for treating subjects having disorders associated either with excessive cell proliferation, such as in the treatment of various cancers, or with aging (degenerative diseases such as Alzheimer and Huntington).
References :
Coste, F., Ober, M., Le Bihan, Y.-V., Izquierdo, M.A., Hervouet, N., Carell, T. and Castaing, B. “Bacterial base excision repair enzyme Fpg recognizes bulky N7-substituted-FapydG lesion via unproductive binding mode.” Chemistry & Biology (2008) 15, 706-717
Illustration: