Prediction of protein structure with the artificial intelligence (AI)-powered program AlphaFold2 – hailed by the Science magazine, the biggest scientific breakthrough in 2021 – has rapidly revolutionised protein science. Trained on a large dataset of experimentally determined protein structures, AlphaFold2 can generate a model of a protein’s three-dimensional (tertiary) structure given its amino-acid sequence (primary structure). AlphaFold2 models are highly reliable, thus offering a good basis for understanding the function of proteins whose experimental structure is not available or is not complete.
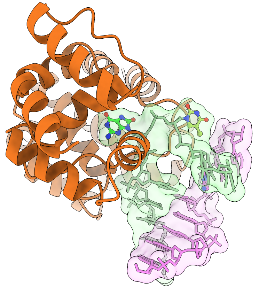
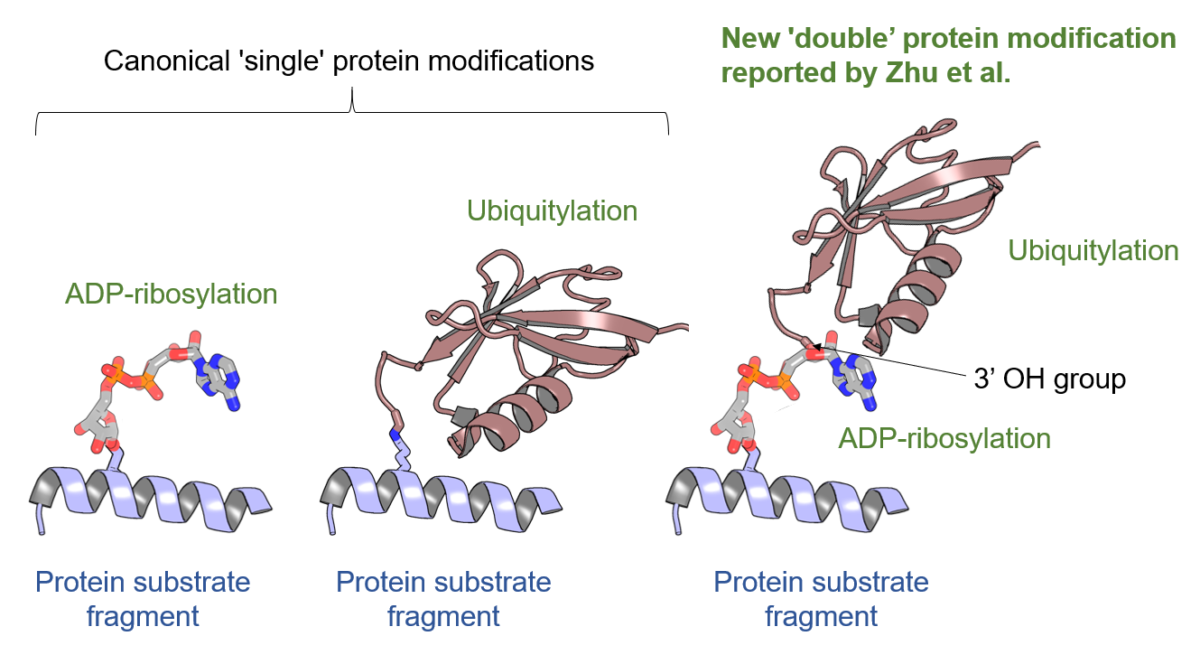
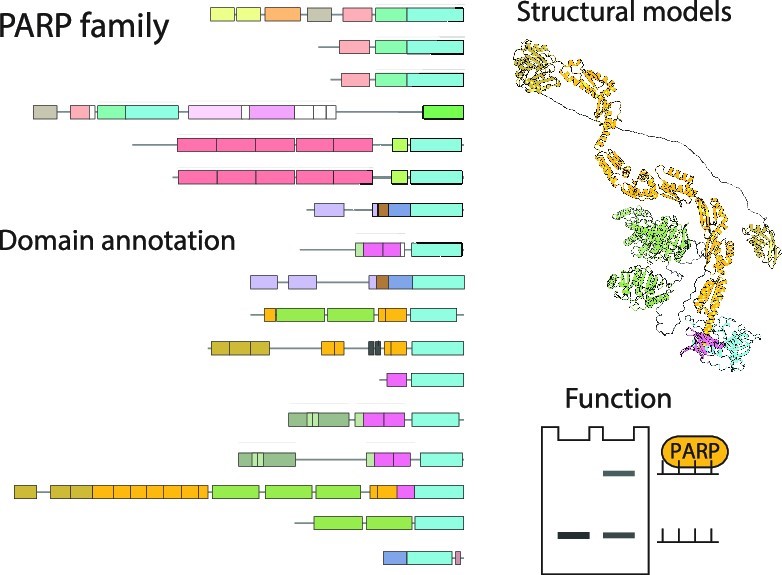
In the present article, published in the journal Nucleic Acids Research, a collaborative team composed of researchers from Orléans, Oxford, and Cambridge, carefully examined AlphaFold2 models of an important group of proteins called the PARP protein family, which includes 17 proteins in human. These proteins regulate DNA repair and many other cellular pathways by catalysing a protein post-translational modification called protein (ADP-ribosyl)ation. The analysis of AlphaFold2 models allowed annotating all protein domains in this family, several of which have not been annotated before. This analysis served as a starting point for various accompanying experiments which validated some of the insights gained from the predicted models. Featuring an accessible introduction into the new computational approaches, the study can serve as a blueprint for scientists studying other protein families.
Two of the CBM members involved in the study are Marcin J. Suskiewicz and Stéphane Goffinont, both from the group “Protein Post-Translational Modifications: Structure, Function, and Dynamics”. This work is linked to a grant from Ligue contre le Cancer CSIRGO 2023.
References :
Marcin J Suskiewicz and others, Updated protein domain annotation of the PARP protein family sheds new light on biological function, Nucleic Acids Research, 2023;, gkad514,
https://doi.org/10.1093/nar/gkad514